Abstract
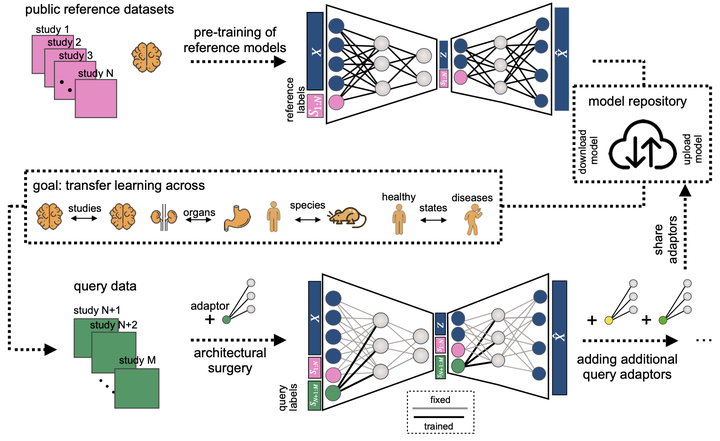
Large single-cell atlases are now routinely generated with the aim of serving as reference to analyse future smaller-scale studies. Yet, learning from reference data is complicated by batch effects between datasets, limited availability of computational resources, and sharing restrictions on raw data. Leveraging advances in machine learning, we propose a deep learning strategy to map query datasets on top of a reference called single-cell architectural surgery (scArches, https://github.com/theislab/scarches). It uses transfer learning and parameter optimization to enable efficient, decentralized, iterative reference building, and the contextualization of new datasets with existing references without sharing raw data. Using examples from mouse brain, pancreas, and whole organism atlases, we showcase that scArches preserves nuanced biological state information while removing batch effects in the data, despite using four orders of magnitude fewer parameters compared to de novo integration. To demonstrate mapping disease variation, we show that scArches preserves detailed COVID-19 disease variation upon reference mapping, enabling discovery of new cell identities that are unseen during training. We envision our method to facilitate collaborative projects by enabling the iterative construction, updating, sharing, and efficient use of reference atlases.